News
Using NSF ACCESS Supercomputers to Improve Tuberculosis Treatment Options
Published May 04, 2026
By Kimberly Mann Bruch
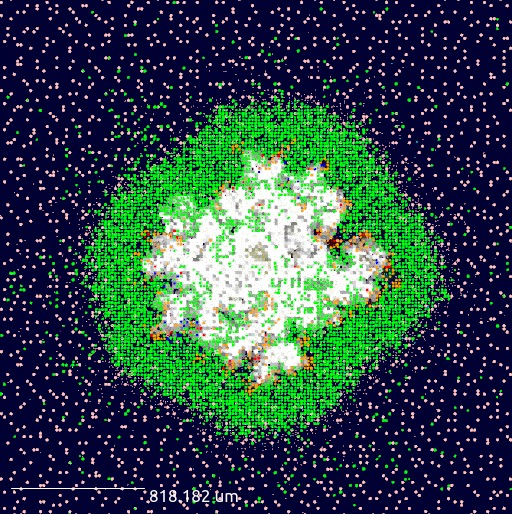
In virtual experiments conducted on SDSC’s Expanse and Purdue’s Anvil, models include tiny infected spheres in the lungs, called granulomas, which form as the immune system tries to control TB bacteria. By studying how drugs reach and affect these granulomas in their simulations, the researchers can better understand which treatment plans are both strong against the infection and easier on patients. Black is lung tissue, green are macrophage cells, white is dead tissue, red squares in black are blood vessels and other colors are different infected cell types. Credit: University of Michigan
A research team from the University of Michigan Medical School, led by mathematical biologist Denise Kirschner, is using U.S. National Science Foundation (NSF) ACCESS allocations on multiple supercomputers to explore better ways to treat tuberculosis (TB). By running computer simulations on machines like Expanse at the UC San Diego Halıcıoğlu School of Data Science and Computing’s San Diego Supercomputer Center (SDSC), they virtually test an array of combinations and doses of antibiotics to see which ones are most likely to work well in patients.
In their latest study, which was published in the Numerical Algebra, Control and Optimization journal, the team created a new computational approach that helps them search through a very large number of possible treatment plans more efficiently. They first run a small set of virtual treatment experiments, then use those results to train a machine learning model that predicts how other, untested treatment plans might perform. A second part of the approach then compares these predicted plans and highlights the ones that appear to offer the best balance between shorter treatment time and lower total drug dose to lower or eliminate bacterial loads.
Using this method, the researchers were able to examine 219 different drug combinations, an unprecedented number for their group and also compared with experimental systems These simulations required more than 600,000 hours of computing time across four NSF ACCESS clusters, including the Expanse system at SDSC and Anvil at the Purdue University Rosen Center for Advanced Computing. The researchers’ results identified treatment plans that, in the virtual experiments, could potentially cure infection in shorter times while using less medicine than some standard regimens, which could help reduce side effects and improve patients’ ability to stay on therapy, especially in areas of the world where drug availability is limited.
“The use of these models generated by SDSC’s Expanse and Purdue’s Anvil helps us work with experimental groups to improve the variety of tuberculosis treatments through different drug doses and drug combinations,” said Kirschner, who is a professor of Microbiology and Immunology at the University of Michigan Medical School. “Without allocations from the NSF to utilize high-performance computing resources, we would not be able to accomplish our work.”
Tuberculosis is a serious, sometimes deadly lung disease caused by bacterial infection and usually requires 6-9 months of antibiotics. Doctors must choose the right mix of drugs and doses for each patient, which can be very challenging. With support from NSF’s ACCESS program, Expanse and Anvil allows the team to simulate how TB bacteria, the immune system and different drugs interact in the body to yield a clearer picture of which treatment options might work best.
The time on Expanse and Anvil was supported by NSF ACCESS (allocation no. MCB140228).

